12k, 24k, 36k…we can now precisely remove over 43k bp from C. elegans genomic DNA!
Knocking out genes is a commonly used tool for biologists to understand gene function in live animals. The examination of phenotypes when a gene is deleted can reveal valuable insights into what role that gene plays in the organism and by extension, in a disease. We have established a CRISPR approach using Cas9 that can be efficiently implemented to create large genomic deletions of at least 36,000 kb.
Why is it a big step?
The genetic manipulation of C. elegans is a crucial approach to investigate genome structural variations and developmental gene regulation. In particular, chromosomal rearrangements such as precise large DNA-fragment deletions, are invaluable tools that will allow researchers to interrogate the role of key genes in human diseases more effectively.
Although RNA interference and loss-of-function mutants have long been used to perturb gene function in C. elegans, both their low efficacy and the off-target effects of RNAi often lead to the need for null mutants that completely eliminate target genes. Such genome editing often requires precise deletion of a large DNA region. Over the last couple of years, the CRISPR-Cas9 system has become the preferred tool for genome engineering, as it enables site-specific alterations in a variety of organisms and cellular contexts. However the use of this technique for the precise deletion of sequences over 24,000 bp has proven to be a major hurdle to overcome in C. elegans.
“[CRISPR/Cas9 technology] typically produces small insertions and deletions (indels) that shift the open reading frame (ORF) of the targeted gene and result in premature termination of translation and loss-of-function phenotypes.” Chen et al 2014.
Why is it hard?
Both the accuracy and efficiency of the processes involved in each of the steps in CRISPR/CAS9 deletion strongly affect the outcome of CRISPR-mediated editing and consequently the utility of the technology. In particular, one concern is the production of an extrachromosomal array created by the portion of DNA that was excised during deletion. Using our proprietary method, we manage to avoid such an issue to create a true, precise deletion of large portions of genomic DNA in C. elegans (see figure 1). To date, we have been able to identify deletion mutants from 13 out of 14 projects.
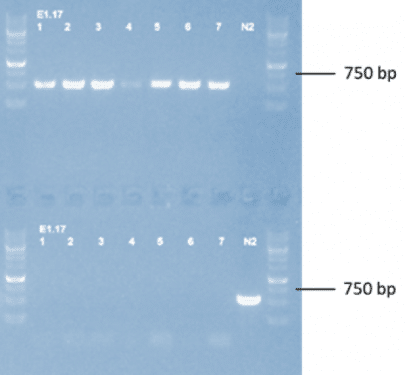
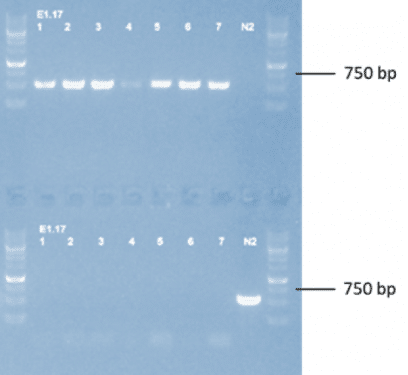
Figure 1: Agarose gel showing genomic DNA from candidate lines amplified by PCR. This gel confirms that the 36,000 bp portion of DNA that was excised during deletion did not create an extrachromosomal array (samples do not contain the DNA that was excised). (Top row) Primers designed to only amplify a band in lines where the gene and unc-2 have successfully been deleted. (Bottom row) Primers designed to only bind in the background strain that still contains the unc-2 gene.
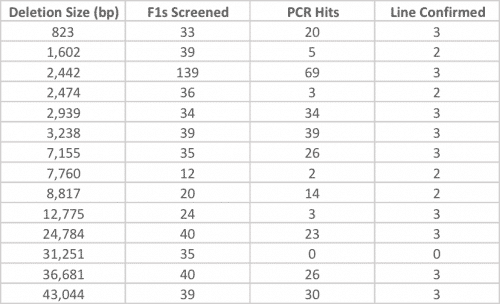
The efficiency of deletion generation does not always directly correlate with deletion size (see Table 1 below).
Videos: Adult C. elegans worms exhibit uncoordinated locomotion pattern after precise deletion of a 35Kb gene. The unc-2 gene, a voltage-gated calcium channel, was deleted resulting in an uncoordinated movement phenotype (video on the bottom) compared with the N2 (video on the top).
N2 (wild-type)
unc-2 KO
We are excited by these results and what it means for the C. elegans research community. Large genomic deletions can now become commonplace and will allow scientists to gain a greater understanding of biology by being able to make complete loss-of-function alleles. Accelerate your research with our top-notch gene editing services. We encourage you to get in touch with us today to explore how our services can help you achieve your research goals.
InVivo Biosystems’ experienced team can take over and deliver the results you need through our C. elegans gene editing services. We welcome the opportunity to discuss your project with you.