Question: What might you get if you cross the feeding organ of a roundworm with a genome wide association study?
Answer: A cornucopia of novel genes for synaptic transmission, neuromodulation, energy regulation, peptidergic signaling, learning and memory, and more.
This riddle, and its answer, are prompted by the convergence of two exciting trends in research on one of biologists’ favorite roundworms, the nematode Caenorhabditis elegans, or C. elegans for short. First, genome wide association analysis (GWA), a powerful new method for gene discovery, has recently been made much easier to do. Second, new methods have come on the scene for measuring the activity of the nematode feeding organ, called the pharynx. And these two trends dovetail beautifully.
That’s because the pharynx is not only a perfect target of GWA studies; it also sits at the intersection of most of the main types of physiological signaling pathways utilized for cellular and neuronal communication in nematodes. Moreover, these are the same pathways found in mammals. So by discovering new genes that correlate with changes in the function of the pharynx, we simultaneously discover new genes in mammals, including ourselves. This kind of research could lead to better understanding of the man human diseases that involve cellular communication, and better drugs to fight them.
But first, more about GWA studies and recording pharyngeal activity.
GWA mapping identifies genes by seeking statistical correlations between numerous multi-genic stretches of DNA on each chromosome and the magnitude of a quantitative phenotype (a continuous variable, like height). In
GWA mapping, the phenotyped entities are individual wild strains from geographically diverse regions. As individuals in a strain are isogenic, the strain’s phenotype can be measured from an almost unlimited number of individuals, greatly increasing statistical power; in human GWA mapping studies, by contrast, the phenotyped entity is a single person, phenotyped once. GWA mapping is unbiased, like a mutagenesis screen, with the advantage that the genetic variants in wild isolates are likely to be relatively benign because highly deleterious alleles have been eliminated by natural selection. This method is therefore immune to the problem of lethal mutations (making some genes hard to discover by mutagenesis), and capable of identifying genes through beneficial aspects of their function.
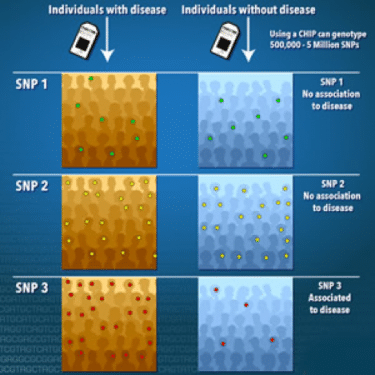
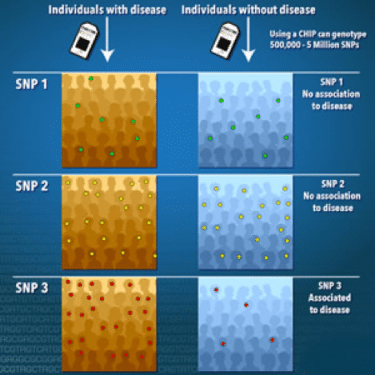 Figure 1. A genome-wide association study is an approach that involves rapidly scanning markers across the complete sets of DNA, or genomes, of many people to find genetic variations associated with a particular disease. Once new genetic associations are identified, researchers can use the information to develop better strategies to detect, treat and prevent the disease. Such studies are particularly useful in finding genetic variations that contribute to common, complex diseases, such as asthma, cancer, diabetes, heart disease and mental illnesses. (source: genome.gov)
Figure 1. A genome-wide association study is an approach that involves rapidly scanning markers across the complete sets of DNA, or genomes, of many people to find genetic variations associated with a particular disease. Once new genetic associations are identified, researchers can use the information to develop better strategies to detect, treat and prevent the disease. Such studies are particularly useful in finding genetic variations that contribute to common, complex diseases, such as asthma, cancer, diabetes, heart disease and mental illnesses. (source: genome.gov)
The accessibility of GWA mapping in C. elegans has recently taken a quantitative leap with establishment of the C. elegans Natural Diversity Resource (CeNDR). CeNDR maintains and distributes more than 600 wild strains, each whole-genome sequenced to a depth of coverage that enables identification of the specific types of variation found at each locus, including single-nucleotide (SNVs), insertion-deletion (indels), transposon insertions, copy-number (CNVs), short-tandem repeats (STRs), genomic rearrangement variants; this depth also enables prediction of a variant’s effects on gene function. Although GWA mapping in C. elegans is a new approach, it has already produced significant results, including identification of sites of resistance to the anthelmintic ivermectin, loci contributing to natural differences in telomere length, and specific mechanisms of chemotherapeutic responses.
Like GWA mapping, recording the activity of the pharynx is now much easier to do. The pharynx is a rhythmically contracting muscular pump which draws food in the form of bacteria particles into the worm’s gut. Its constrictions, like those in the mammalian heart, are generated by large, long lasting electrical pulses called action potentials (Figure 2). Pharyngeal action potentials can be recorded by attaching electrodes to the worm just as cardiac action potentials can be recorded by attaching EKG electrodes to a patient. This used to be hard to do because C. elegans is tiny. Each one looks like a piece of lint, or a comma in 7 point font. But now, thanks to an approach called softlithograpy microfluidics, it’s possible to make electrophysiology “chips” containing microscopic, fluid-filled channels that worms can be pumping into. Electrodes attached to the channels pick up the action potentials which are recorded like blips on a heart monitor and stored on a computer for later analysis. Sucker recordings are called electropharyngeograms, or EPGs. Using this new approach, it’s easy to record EPGs from many tens of worms per hour.
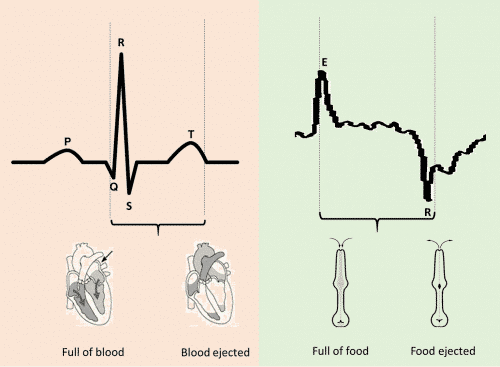 Figure 2. The cardiac and pharyngeal pumping behaviors share similarities which can be exploited and explored using the ScreenChip phenotyping platform.
Figure 2. The cardiac and pharyngeal pumping behaviors share similarities which can be exploited and explored using the ScreenChip phenotyping platform.
The reason the GWA and the pharynx go together so well is that the EPGs can be quantified in numerous ways. In fact, an almost unlimited number of quantitative phenotypes is hidden in each EPG recording. For example, there are the basic measures of electrical activity such as average action-potential frequency, duration, and amplitude. And for every averaged quantity, there is its variance, a measure of event-to-event variability. Going further, one can utilize higher-order statistical methods such histograms, which are graphs whose shapes quantify how individual measures are distributed above and below their average. For example, a histogram can quantify the extent to which this distribution is symmetrical or skewed.
But the big message here is that each quantitative phenotype is the basis of a unique GWA map, and each mapping shines a spotlight on a particular subset of genes. There will almost certainly be partial overlap between these subsets, but each phenotype will likely be associated with some, possibly many genes that do not appear in any other subsets. Thus, a single collection of EPG recordings simultaneously probes a large number of unique regions of the genome, a number limited only by one’s imagination in constructing quantitative phenotypes. In this way, the genetic cornucopia is likely to fill quickly.